Note
Go to the end to download the full example code.
Reblocking#
import tempfile
import pandas as pd
from pathlib import Path
import pyvista as pv
from parq_blockmodel import ParquetBlockModel
Create a Parquet Block Model#
We leverage the create_demo_block_model class method to create a Parquet Block Model.
temp_dir = Path(tempfile.gettempdir()) / "block_model_example"
temp_dir.mkdir(parents=True, exist_ok=True)
pbm: ParquetBlockModel = ParquetBlockModel.create_demo_block_model(
filename=temp_dir / "demo_block_model.parquet")
pbm
ParquetBlockModel(name=demo_block_model, path=/tmp/block_model_example/demo_block_model.pbm)
Visualise the Model#
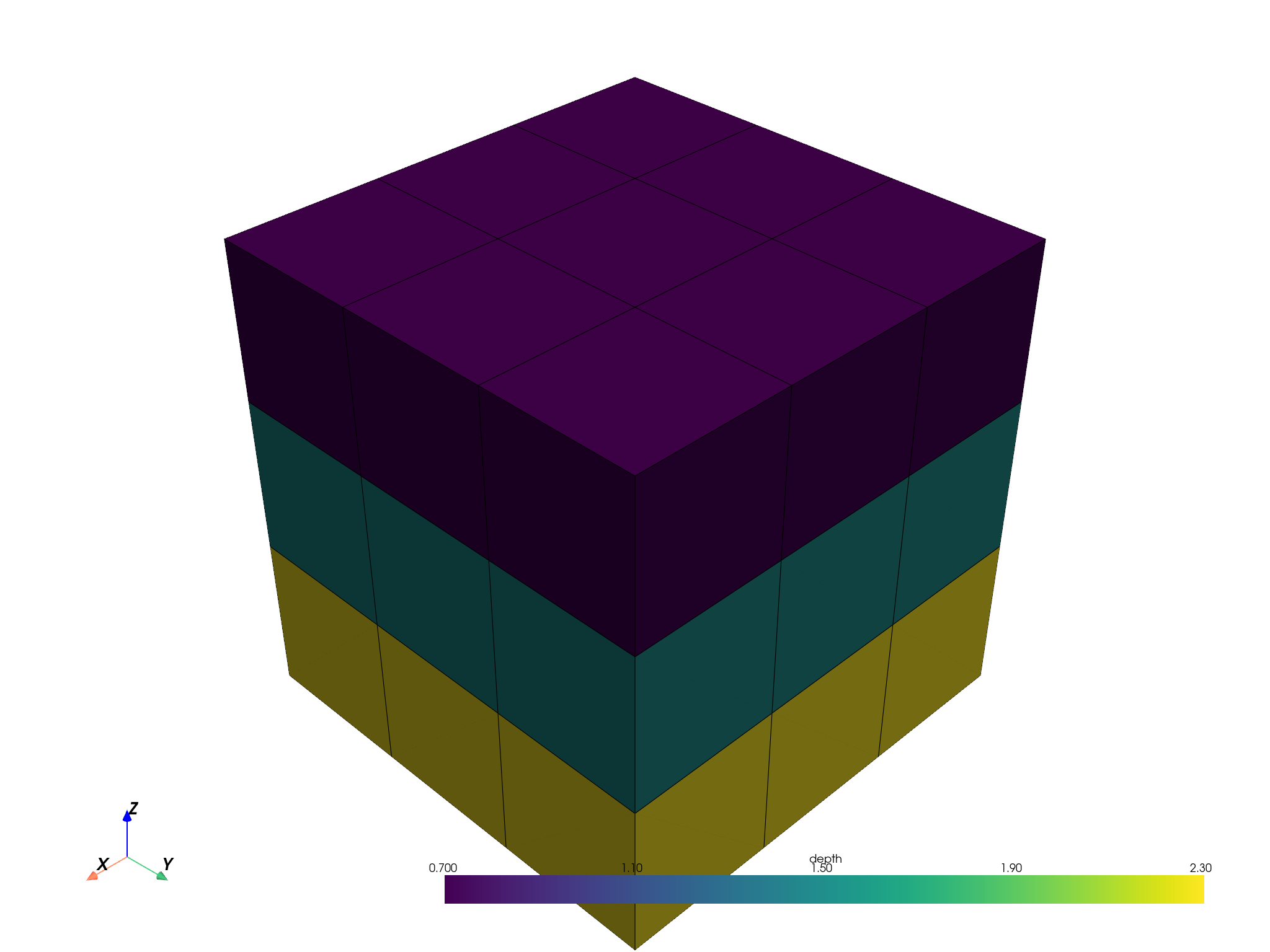
A continuous attribute.
pbm.plot(scalar='depth', threshold=False, enable_picking=True).show()
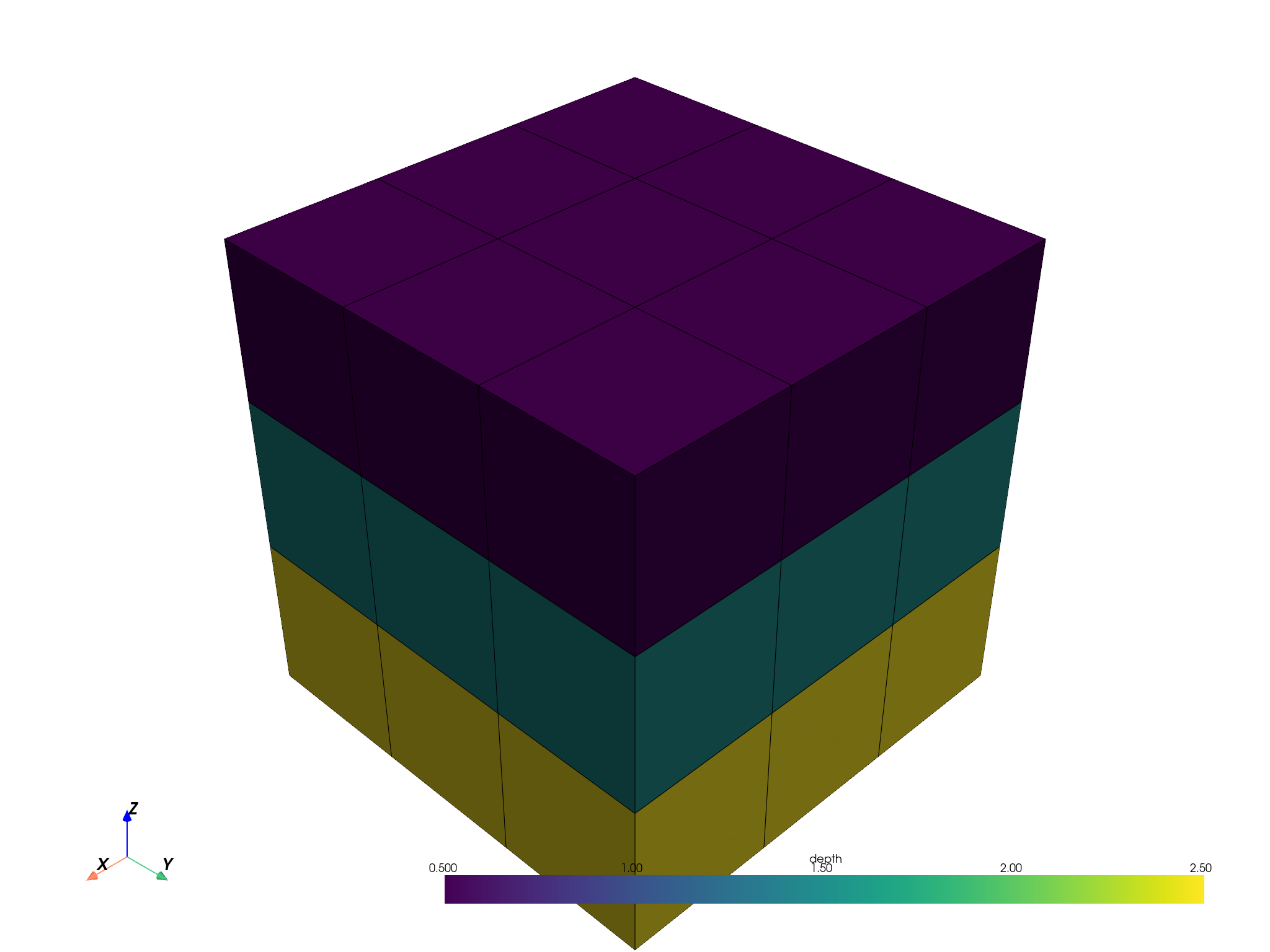
A categorical attribute.
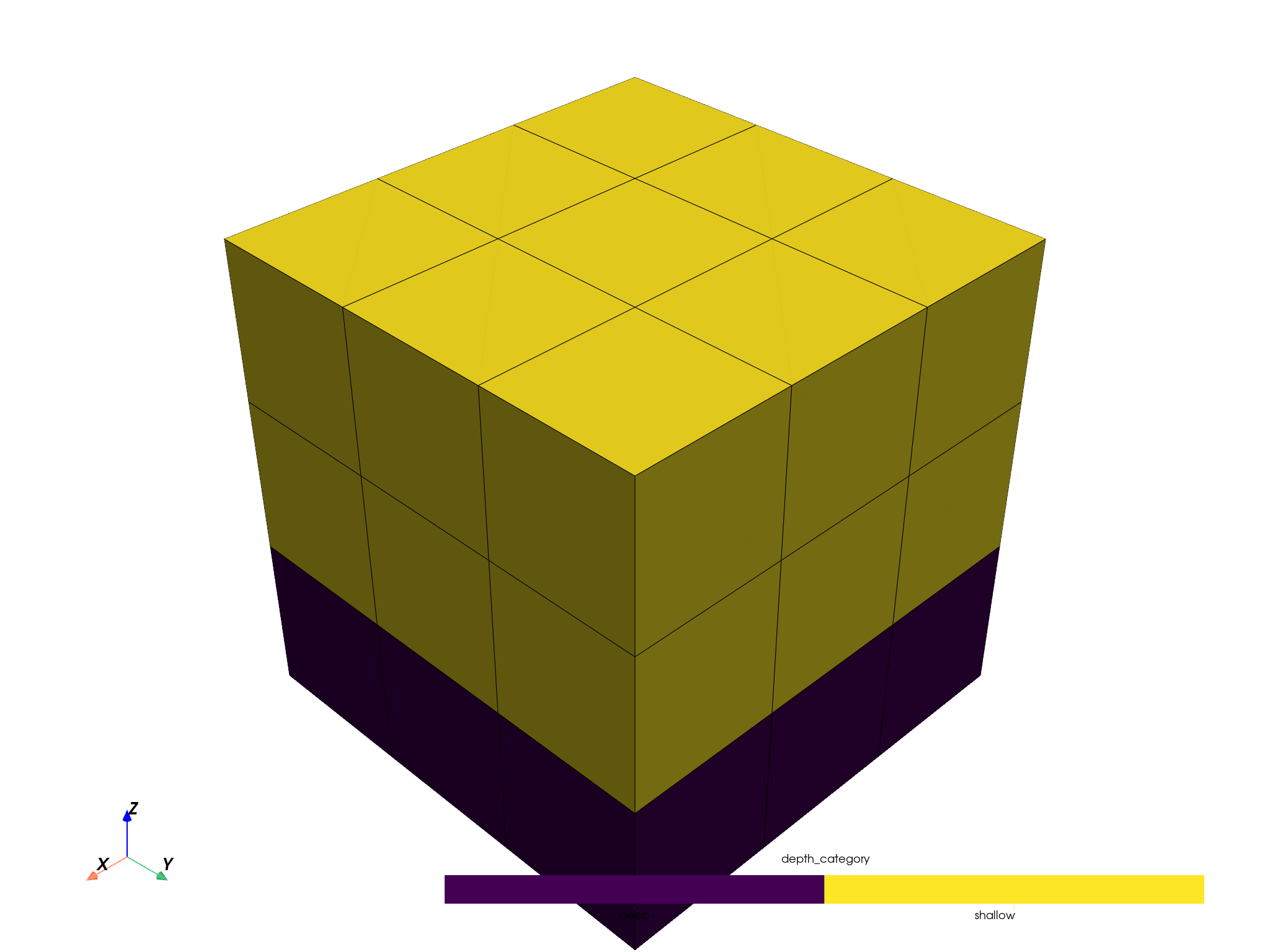
pbm.plot(scalar='depth_category', threshold=False, enable_picking=True).show()

Upsampling#
Create a new grid with smaller blocks, and visualise.
upsampled_pbm: ParquetBlockModel = pbm.upsample(new_block_size=(0.5, 0.5, 0.5),
interpolation_config={
'depth': 'linear',
'depth_category': 'nearest'
})
upsampled_pbm.plot(scalar='depth', threshold=False, enable_picking=True).show()
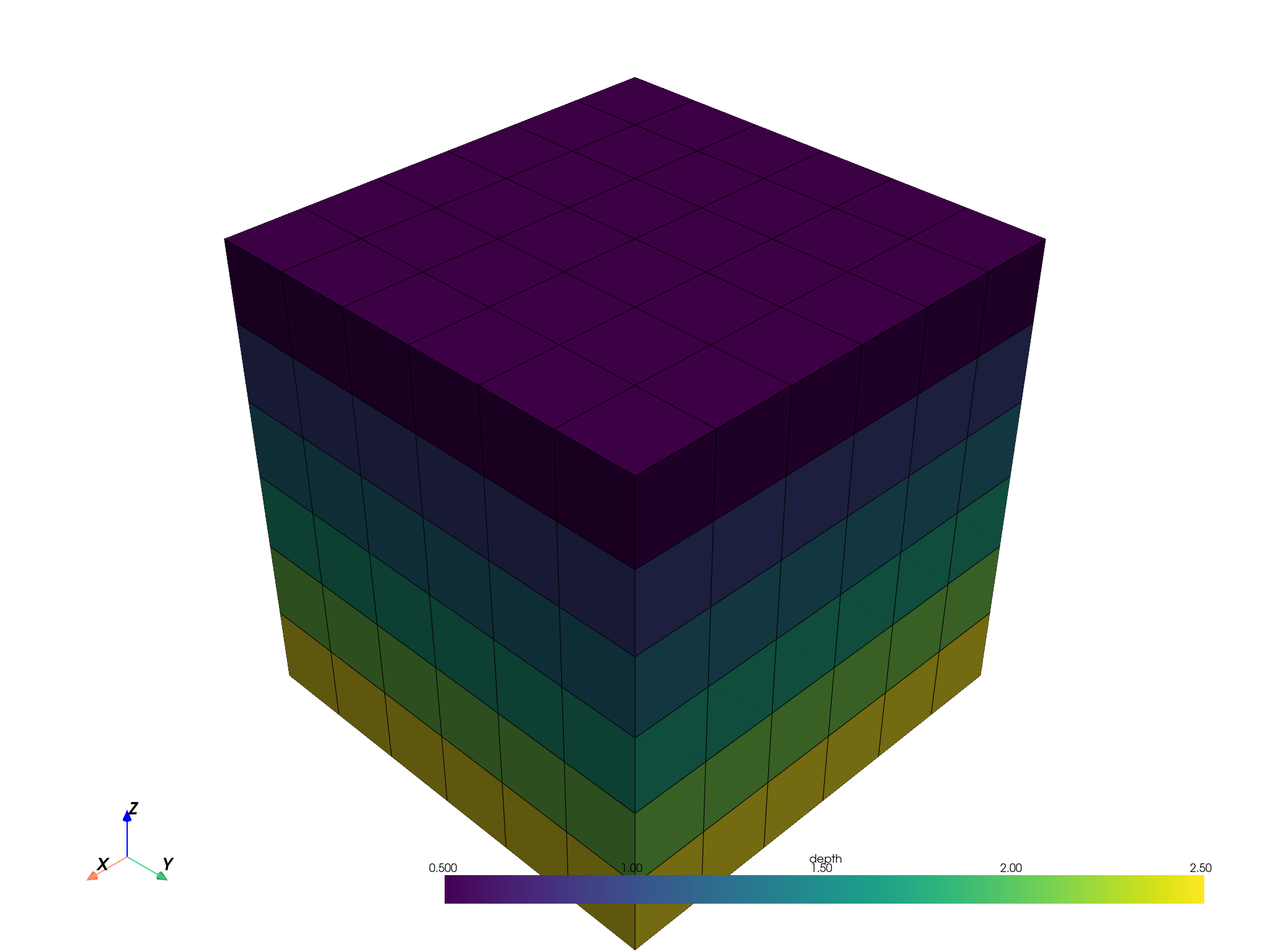
Visualise a reblocked categorical attribute.
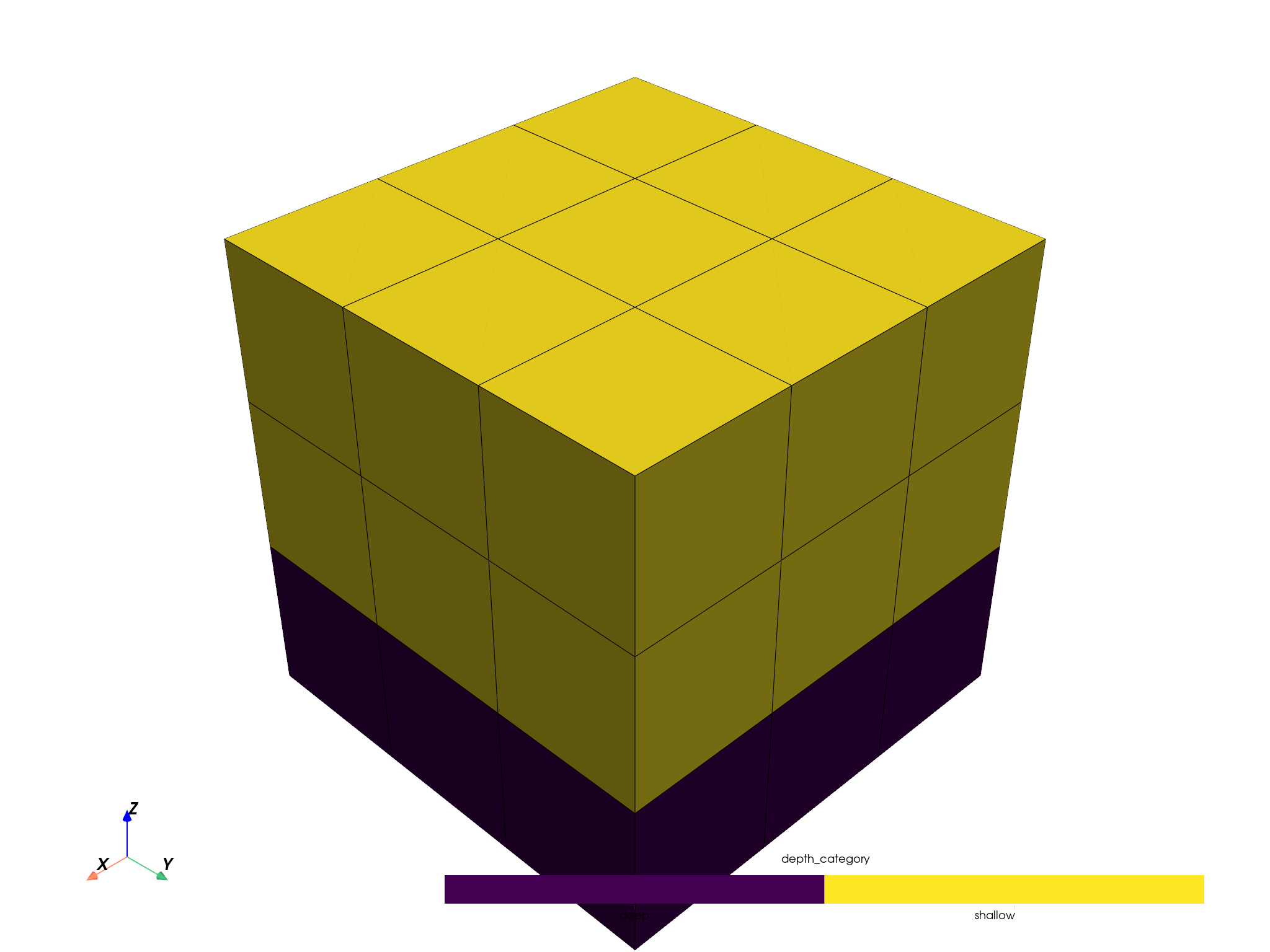
upsampled_pbm.plot(scalar='depth_category', threshold=False, enable_picking=True).show()

Downsample#
Downsample the upsampled model back to the original block size.
downsampled_pbm: ParquetBlockModel = upsampled_pbm.downsample(new_block_size=(1.0, 1.0, 1.0),
aggregation_config={
'depth': {'method': 'mean'},
'depth_category': {'method': 'mode'}
})
downsampled_pbm.plot(scalar='depth', threshold=False, enable_picking=True).show()
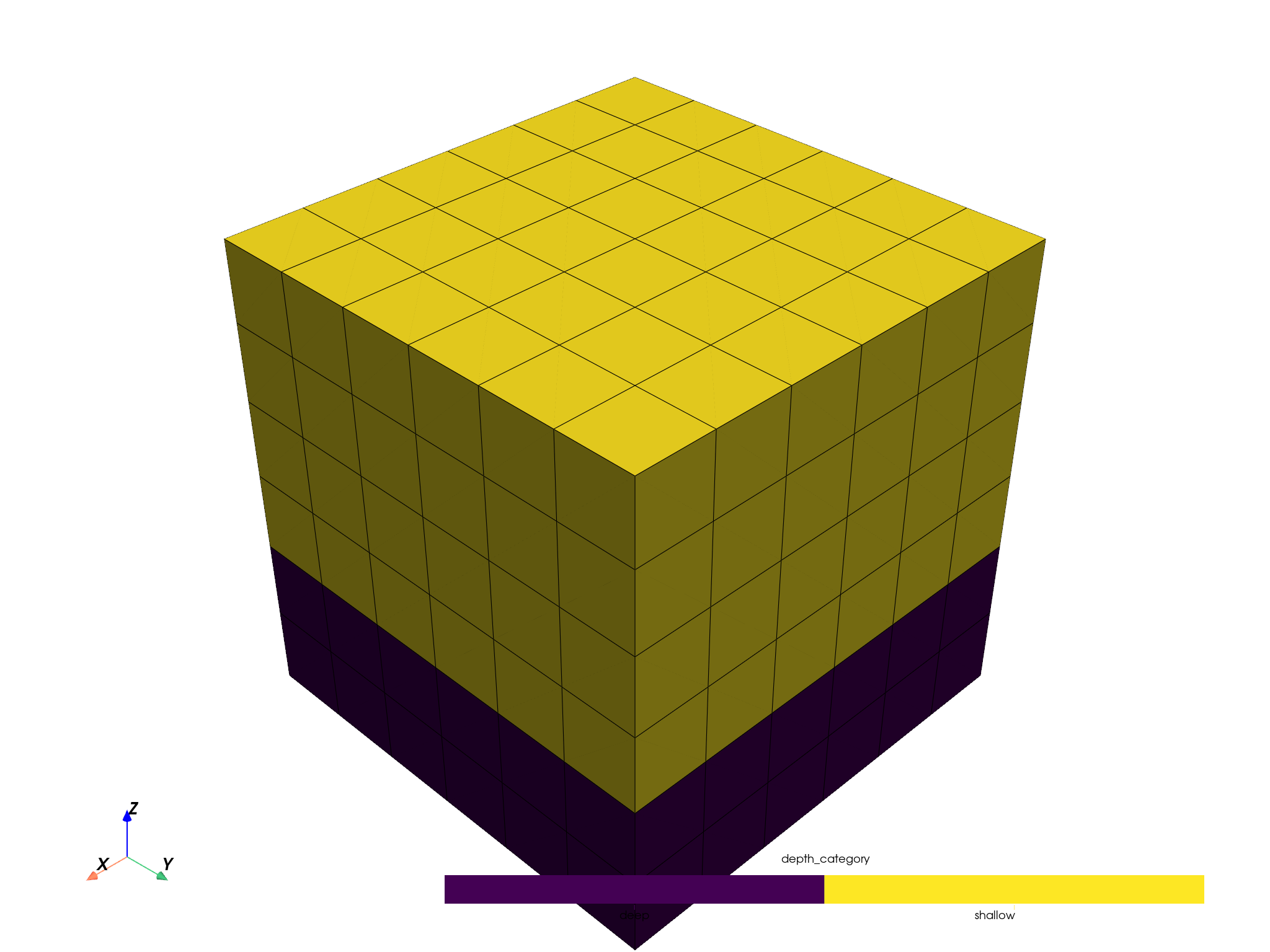
downsampled_pbm.plot(scalar='depth_category', threshold=False, enable_picking=True).show()
Validate#
"""Validate that reblocking preserves both data and centroids.
1. Confirm the round trip (upsample then downsample) returns the
original *data* when viewed in a consistent ijk ordering.
2. Demonstrate that centroids derived from geometry are as expected.
"""
# Read attribute data on the canonical ijk grid for both models. We work
# only with the attributes of interest here (depth and depth_category)
# rather than asserting equality for every auxiliary column.
blocks_original: pd.DataFrame = pbm.read(index="ijk", dense=True)[["depth", "depth_category"]]
blocks_up_down: pd.DataFrame = downsampled_pbm.read(index="ijk", dense=True)[["depth", "depth_category"]]
# 1a. Ensure the reblocked attributes are fully populated (no NaNs) on
# the dense ijk grid for this dense demo model.
for col in ["depth", "depth_category"]:
assert blocks_up_down[col].notna().all(), f"{col} contains NaNs after reblocking"
# 1b. Numeric attribute: values should be similar after an
# upsample→downsample round trip, but not necessarily identical. Check
# that the maximum absolute difference is within a small tolerance.
depth_diff = (blocks_up_down["depth"].sort_index() -
blocks_original["depth"].sort_index()).abs()
assert depth_diff.max() < 0.5, (
"Depth changed too much during reblocking; max |Δdepth| = "
f"{float(depth_diff.max()):.3f}"
)
# 1c. Categorical attribute: in this demo we expect depth_category
# labels to be preserved exactly by the round trip. We compare the
# string representations to ignore differences in categorical metadata
# such as the "ordered" flag.
orig_cat = blocks_original["depth_category"].astype(str).sort_index()
down_cat = blocks_up_down["depth_category"].astype(str).sort_index()
pd.testing.assert_series_equal(orig_cat, down_cat, check_names=True)
# Demonstrate that the logical ijk grid and XYZ centroids of the
# downsampled model match those of the original geometry.
orig_ijk = pbm.geometry.to_multi_index_ijk()
down_ijk = downsampled_pbm.geometry.to_multi_index_ijk()
pd.testing.assert_index_equal(orig_ijk, down_ijk)
orig_xyz = pbm.geometry.to_multi_index_xyz()
down_xyz = downsampled_pbm.geometry.to_multi_index_xyz()
pd.testing.assert_index_equal(orig_xyz, down_xyz)
Total running time of the script: (0 minutes 2.451 seconds)